Classification of viruses on the basis of genome
Introduction
An ongoing effort to classify viruses similar to that for cellular organisms has been complicated by the fact that viruses evolve rapidly and they can transfer their genes between viruses and with cellular organisms.
Taxonomic scheme for all Viruses Has yet to Be Universally Adopted
- Viral nomenclature has used a variety of virion features.
- The measles virus and poxviruses, for example, are named after the disease they cause; the Ebola and Marburg viruses after the location from which they were originally isolated; and the Epstein-Barr virus after the researchers who studied it.
- Other viruses are named after morphological factors—the coronaviruses (corona = “crown”) have a crown-like capsid, and the picornaviruses (pico = “small”; rna = “ribonucleic acid”) are very small viruses with an RNA
genome. - Such characteristics, however, do not reflect the evolutionary history of the viruses.
- A more encompassing classification system based on evolutionary relationships is being devised by the International Committee on Taxonomy of Viruses (ICTV).
- At this writing, higher-order taxa (phyla and classes) have not been completely developed.
- To date, six orders are recognized that comprise 87 families, each ending with viridae (e.g., Herpesviridae).
- However, many other viruses have not yet been assigned to a family.
- Viruses have been categorized into hundreds of genera; each genus name ends with the suffix -virus (e.g., human herpes virus).
- In this text, we mostly use the family name (e.g., Herpesviridae) when referring to the whole family of viruses and the common name (e.g., herpes simplex virus) when discussing a specific virus in a family.
- Animal viruses usually are classified into two groups based on their genome (DNA or RNA) and then split into separate families based on characteristics such as strand type (double stranded and single stranded) and presence or absence of an envelope.
DNA Viruses classification
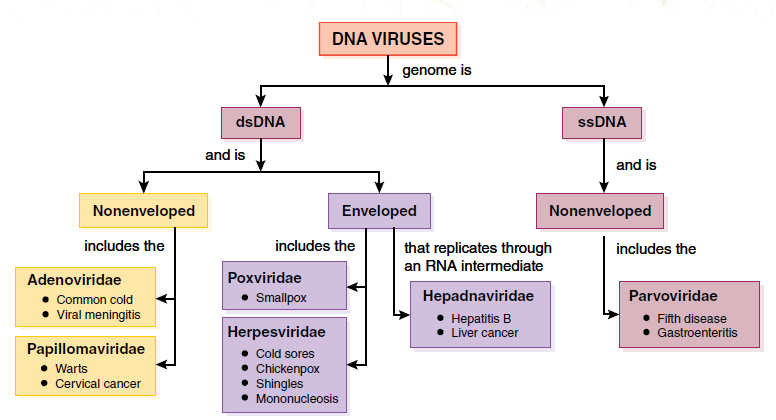
- Many viruses contain either double-stranded (ds) or single-stranded (ss) DNA genomes with nucleocapsids that are nonenveloped or enveloped.
- Among the nonenveloped dsDNA viruses are the Adenoviridae, which includes viruses causing common colds, and the Papillomaviridae, which includes members causing common skin warts.
- The enveloped dsDNA viruses include the Herpesviridae whose members include viruses responsible for cold sores (fever blisters) and chickenpox; the Poxviridae that includes the smallpox virus; and the Hepadnaviridae that includes the hepatitis B virus.
- The ssDNA viruses are nonenveloped.
- The Parvoviridae include one virus that causes a childhood rash called fifth disease.
- A separate canine parvovirus infects the lymph nodes and intestines of puppies.
RNA Viruses classification
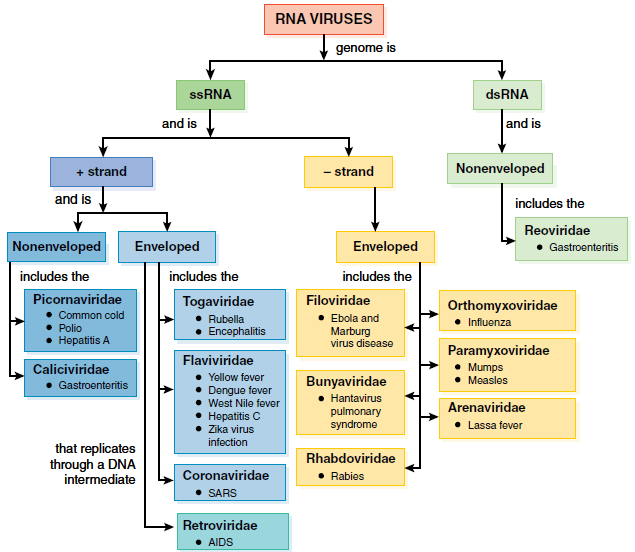
- A large number of viruses have RNA genomes consisting of either dsRNA or ssRNA.
- The viruses that have dsRNA include the nonenveloped Reoviridae, whose genomes are segmented and surrounded by a two-layered capsid. In this family, the rotaviruses can cause severe diarrhea and intestinal distress (gastroenteritis) in infants and young children.
- The ssRNA viruses have nucleocapsids that are enveloped or nonenveloped.
- Their genomes are classified according to the sense or polarity of their RNA into positive sense and negative sense.
- Please refer to this figure as we describe the different groups of viruses by their type of genome.
Positive-Sense (+sense) rna Viruses
- The ssRNA in the +sense RNA viruses acts as mRNA and, on infection of a host cell, the RNA can be translated directly by the host cell ribosomes (thus called + sense).
- The nonenveloped Picornaviridae contain viruses responsible for many well-known human diseases, including common colds, polio, and hepatitis A.
- The enveloped families, such as the Flaviviridae, also include many viruses responsible for human diseases, such as yellow fever, West Nile disease, hepatitis C, and Zika virus infection. In fact, the +sense RNA viruses are the largest group of RNA viruses.
- Although the Retroviridae are singlestranded, +sense RNA viruses, this family of viruses behaves in an unusual way.
- After infecting a host cell, the virus uses its own enzyme to produce dsDNA from its ssRNA genome through a process of reverse transcription (retro = “backwards”).
- Only after the new dsDNA has integrated into the host cell genome can the viral DNA genes be transcribed into mRNA and translated.
- The human immunodeficiency virus (HIV) that is responsible for AIDS is the most recognized virus in this family.
Negative-sense (–Sense) rna Viruses
- The ssRNA in –sense RNA viruses is complementary in base sequence to mRNA (thus called – sense).
- As a result, the RNA must be made into a “readable” form.
- This is accomplished when a RNA polymerase converts the –sense RNA into a +sense RNA prior to translation. The +sense RNA then is translated by host ribosomes.
- The virus families in this group are enveloped, and several members are the infectious agents for a considerable number of human diseases, including Ebola virus disease, influenza, measles, mumps, and rabies.
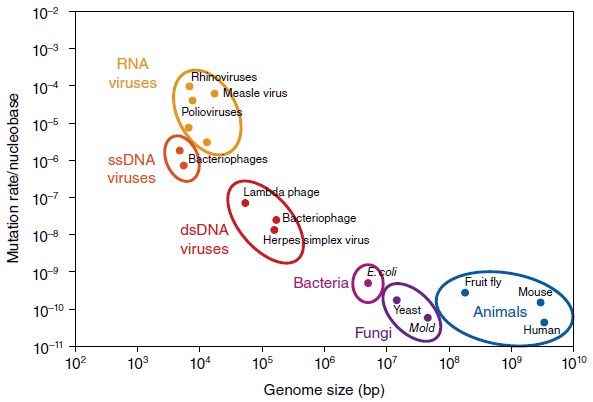
As a rule, the genomes of RNA viruses are smaller than those in DNA viruses, and they depend more heavily on host cell proteins and enzymes for replication. RNA virus genomes also are more error (mutation) prone when copying their RNA genome because the RNA polymerase lacks the efficient proofreading exhibited by DNA polymerase to correct replication errors.
Thus, RNA viruses, such as the influenza viruses, tend to “genetically drift,” evolving more rapidly into new, potentially epidemic strains.
Reference and Sources
- https://www.researchgate.net/publication/24444547_Nudiviruses_and_other_large_doublestranded_circular_DNA_viruses_of_invertebrates_New_insights_on_an_old_topic
- https://www.researchgate.net/publication/334496812_Evolution_and_ecology_of_plant_viruses
- https://quizlet.com/198265167/chapter-14-the-viruses-and-virus-like-agents-flash-cards/
- https://quizlet.com/514275106/microbiology-exam-2-flash-cards/
- https://quizlet.com/99275101/biol-1353-chapter-13-multiple-choice-quiz-flash-cards/
- http://www.microbiologybook.org/mhunt/rna-ho.htm
- https://quizlet.com/143573017/microbiology-chapter-27-flash-cards/
Also Read:
- What is Gene Expression?
- Microbial Fuel Cells
- what is microbiology?
- Animal and plant viruses, prions, and viroids
- Text and Practical Microbiology for MLT
- How to Build a Microorganism?
- Different types of Pathways for ATP Production
- Difference between Prokaryotes and Eukaryotes
- Roles of Viruses In Aquatic Ecosystems
- Bacteriophage: characteristics and replication of lytic and lysogenic cycle
- Bacterial Growth Curve: Definition, Phases and Measurement
- Probiotics: Introduction, Development and Uses in Agriculture
- Cider: Production, Extraction, Fermentation and Maturation
- Milk: Composition, Processing, Pasteurization, Pathogens and Spoilage
- Chromatography: Introduction, Principle, Classification and applications
