Introduction
“Building a microorganism” means designing and assembling a living cell—typically a bacterium—from scratch or by modifying existing genetic material. This is the realm of synthetic biology, a multidisciplinary field that combines biology, genetics, engineering, and computer science to design and construct new biological systems.
- Biotechnology has repeatedly demonstrated how useful the genetic modification of an organism can be.
- From the production of recombinant human insulin to recombinant indigo (the dye used in blue jeans), it is clear that “gene juggling” can potentially be used for the production of a wide range of products.
But what if, rather than modifying one gene at a time, it were possible to design the ideal genome?
- The construction of such a genome would enable synthesis of the precise genotype best suited for the application at hand, whether that is generating biofuels or pharmaceuticals.
The first step in reaching this goal was to build a genome from scratch.
- Molecular biologists at the J. Craig Venter Institute (JCVI) accomplished this in 2008 with the one of the smallest bacterial genomes, that of Mycoplasma genitalium.
- The assembly of chemically synthesized DNA was performed in a series of BACs and then YACs to generate the entire bacterial chromosome in a yeast host.
- A year later, JCVI scientists showed it was possible to “transplant” a bacterial genome from a yeast host into a bacterium.
- So by 2010 the team was ready to go the distance and construct a synthetic genome and transplant it into a bacterial host.
- The JCVI researchers used M. mycoides as their model genome.
- At 1.1 million base pairs (Mb), this microbe’s genome is about twice the size of M. genitalium.
- The first step was to chemically synthesize 1,078 oligonucleotides, each 1080 bp long.
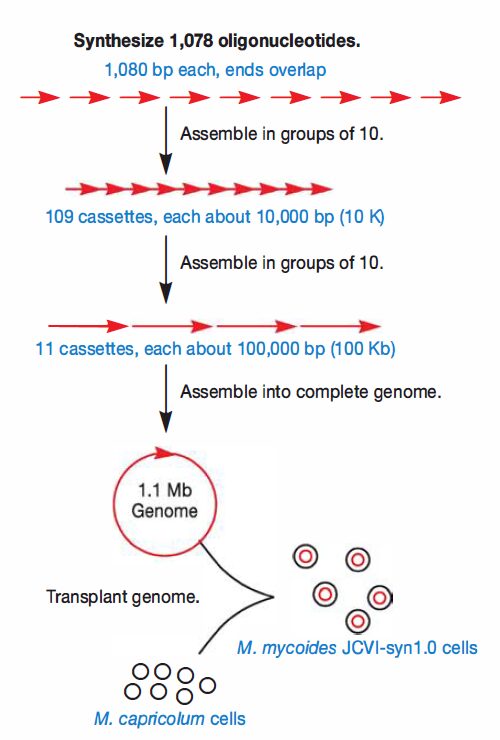
- Importantly, 80 bp at each end of every oligonucleotide overlapped with its neighbor, so within an E. coli host, these could be assembled in the same order as that found in the native chromosome.
- Assembly in groups of 10 generated 109 chromosomal fragments, each about 10,000 bp.
- These were then transformed into yeast, where they were joined into 11 larger cassettes of 100,000 bp each. Finally, these were recombined to yield the entire genome.
- In addition to the native genes, four nucleotide sequences, called watermarks, were included.
- Just as a watermark is used to identify the origin of a piece of stationary or currency, these nucleotide sequence watermarks were added to identify the genome as artificial.
- The final step was to transplant the synthetic genome from the yeast host to the final bacterial host, M. capricolum.
- Just as in any other cloning experiment, this depended on the presence of a selectable marker. In this case, a gene that conferred resistance to the antibiotic tetracycline was included in the synthetic genome.
- Upon selection of tetracycline-resistant M. capricolum cells bearing the M. mycoides genome, a series of verification tests were performed.
- These involved PCR and analysis of restriction endonuclease digestion patterns.
- Once it was confirmed that the host cell was carrying the synthetic genome, all that was left to do was to name the new organism. The JCVI scientists dubbed their new organism M. mycoides JCVI-synl.O.
Reference and Sources
- https://science.sciencemag.org/content/328/5981/news-summaries.full
- https://www.researchgate.net/publication/23273981_Microbial_Biosynthesis_and_Biotransformation_of_Indigo_and_Indigo-like_Pigments
- https://www.researchgate.net/publication/12050315_Cloning_and_Expression_of_a_Ralstonia_eutropha_HF39_Gene_Mediating_Indigo_Formation_inEscherichia_coli
Also read:
- Whole-Genome Shotgun Sequencing: overview, steps and achivements
- Cyanobacteria: occurrence, morphology, structure, reproduction
- Downstream processing and its steps
- Streptomycin : chemical structure, production , recovery and uses.
- Nosocomial Infection: Introduction, Source, Control and Prevention
- Mesothelioma Lung Cancer
- Dodder: Introduction, Development of Disease, Symptoms and Control
- Virus: Introduction, Properties and Classifications
- Animal and plant viruses, prions, and viroids

