Central Dogma of Molecular Biology
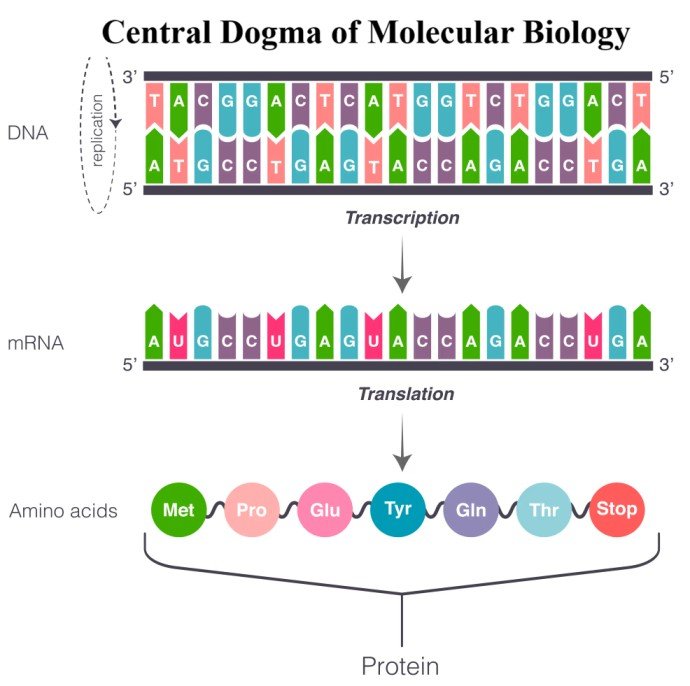
The Central Dogma of Molecular Biology explains how genetic information flows inside a biological system. Proposed by Francis Crick in 1958, this principle describes the transfer of genetic information from DNA → RNA → Protein.
According to the central dogma, the genetic instructions stored in DNA (Deoxyribonucleic Acid) are first copied into RNA (Ribonucleic Acid) and then translated into proteins, which perform most of the structural and functional roles in living organisms.
In simple terms: DNA → RNA → Protein
This flow of genetic information controls the development, functioning, and reproduction of all living organisms.
The central dogma involves three major molecular processes:
- Replication – DNA makes an identical copy of itself
- Transcription – DNA is converted into RNA
- Translation – RNA is converted into proteins
These processes ensure that genetic information is accurately transmitted and expressed in cells.
1. DNA Replication
Definition
DNA replication is the biological process by which a cell produces an identical copy of its DNA before cell division.
Replication is described as semiconservative replication, meaning each new DNA molecule contains:
- One parental strand
- One newly synthesized strand
This mechanism ensures accurate inheritance of genetic information from one generation of cells to the next.
Replication occurs during the S phase of the cell cycle in eukaryotic cells.
Mechanism of DNA Replication
DNA replication occurs in three main stages:
- Initiation
- Elongation
- Termination

Initiation
- In the genome, the origin of replication is a string of base pairs at which DNA replication starts; these sequences are typically rich in AT content, which facilitates separation.
- The weakest bonds are those of at bonds since they contain the fewest hydrogen bonds. Unlike prokaryotes, which have a single origin of replication, eukaryotes have many.
- The process of replication starts when origin-binding proteins attach to the DNA’s origin of replication.
- The enzyme helicase then initiates the unwinding of the double helix, resulting in a replication fork.
- The leading and lagging strands are created at the Y-shaped replication fork. By breaking hydrogen bonds, helicase initiates unwinding.
- The unwound DNA is kept stable by single-stranded binding proteins (SSBs), which prevent it from folding into secondary structures.
- These secondary structures might block the
Elongation
- Topoisomerases alleviate supercoiling as DNA unwinds (Topoisomerase I relax, Topoisomerase II/DNA gyrase introduces negative supercoils in prokaryotes). To provide a starting point, primase deposits brief RNA primers.
- DNA polymerase III in prokaryotes generates DNA in the 5’→3′ orientation with 3’→5′ proofreading. In eukaryotes, DNA polymerase α initiates replication before passing it over to polymerase ε (leading strand) and polymerase δ (lagging strand).
- The primary strand is made in a continuous manner in the direction of the replication fork. Each Okazaki piece of the lagging strand requires a primer since it is produced in a discontinuous manner. The RNA primers are replaced by DNA-by-DNA
Termination
- The procedure of extending the new DNA strands continues until either the DNA template strand has nothing more to replicate (i.e., at the end of the chromosome) or two replication forks converge and then stop.
- The interaction of two replication forks is uncontrolled and occurs at random throughout the chromosome. The freshly created strands are linked and stabilized after DNA synthesis has been completed.
- Two enzymes are necessary to accomplish this stabilization for the lagging strand: DNA ligase joins the Okazaki fragments together to form a single continuous strand, and RNAase H removes the RNA primer at the start of each fragment.
Functions of DNA Replication
DNA replication is essential for life because it:
- Ensures accurate genetic inheritance
- Allows cell growth and division
- Maintains genomic stability
- Enables proper gene expression
Without replication, cells could not divide or transmit genetic information to daughter cells.
2. Transcription
Definition
- Transcription is the process by which genetic information stored in DNA is copied into messenger RNA (mRNA).
- The enzyme responsible for transcription is RNA polymerase.
- This process mainly occurs in the nucleus of eukaryotic cells and in the cytoplasm of prokaryotes.
- The synthesized mRNA carries genetic instructions from DNA to the ribosome where proteins are produced.
Mechanism of Transcription
Transcription occurs in three main steps:
- Initiation
- Elongation
- Termination

Initiation
- The enzyme RNA polymerase, which binds to and travels along the DNA molecule until it identifies a promoter sequence, initiates transcription.
- The transcription start site is indicated by this region of DNA, and a DNA molecule may contain many promoter sequences.
- Transcription factors are proteins that regulate the pace of transcription. They also bind to the promoter sequences along with RNA polymerase.
- RNA polymerase unwinds a portion of the DNA double helix, revealing the bases on each of the two DNA strands after it has attached to the promoter sequence.
Elongation
- One DNA strand, the template strand, reads in a 3′ to 5′ (three-prime to five-prime) orientation and serves as the template for the novel mRNA molecule. The coding strand is the name of the other DNA strand.
- This is due to the fact that its base sequence is the same as the produced mRNA, with the exception of the substitution of thiamine bases for uracil.
- Incoming ribonucleotides are used by RNA polymerase to create the new mRNA strand.
- It accomplishes this by utilizing complementary base pairing (A to U, T to A, C to G, and G to C) to catalyze the creation of phosphodiester linkages between neighboring ribonucleotides. Since bases can only be inserted to the 3′ end, the strand grows in a 5′ to 3′ direction.
Termination
- Transcription ends when RNA polymerase reaches a termination sequence.
- There are two major types of termination in prokaryotes.
- Rho-Dependent Termination
- Rho-Independent Termination
Rho-Dependent Termination
- It requires Rho protein, an ATP-dependent helicase.
Steps:
- Rho binds to RNA
- Moves along RNA transcript
- Unwinds RNA-DNA hybrid
- Releases RNA transcript
Rho-Independent Termination
- Does not require Rho protein.
Characteristics:
- GC-rich inverted repeat sequence.
- Formation of hairpin loop in RNA.
- Followed by poly-U sequence.
This weakens RNA-DNA binding and releases RNA polymerase.
Post-Transcriptional Modifications (Eukaryotes)
Eukaryotic mRNA undergoes several modifications:
- 5′ Cap- Added to protect mRNA and assist ribosome binding.
- Poly-A Tail- Added to the 3′ end for stability.
RNA Splicing- Removal of introns and joining of exons by the spliceosome. These modifications produce mature mRNA ready for translation.
Enzymes & Their Role
| Enzyme / Factor | Function | Prokaryotes | Eukaryotes |
| RNA Polymerase | Synthesizes RNA from DNA template (5′→3′) | Single RNA polymerase (σ factor helps initiation) | Three types: I (rRNA), II (mRNA, snRNA, miRNA), III (tRNA, 5S rRNA) |
| Helicase / TFs | Unwinds DNA at promoter region | Helicase helps unwind | General transcription factors (e.g., TFIID binds TATA box) assist unwinding |
| Topoisomerase | Relieves supercoiling ahead of transcription bubble | Present | Present |
| Capping Enzymes | Add 5′ cap for stability and ribosome recognition | does not occur | Present |
| Spliceosome (snRNPs) | Removes introns, joins exons in pre-mRNA | does not occur | Present |
| Poly(A) Polymerase | Adds poly-A tail at 3′ end of mRNA | does not occur | Present |
Function
- The transmission of genetic data Transcription creates an RNA strand (mainly mRNA) from the DNA sequence, which functions as a messenger transporting genetic information from the nucleus (or nucleoid in prokaryotes) to the location of protein production.
- The synthesis of functional RNAs – It produces various forms of RNA (mRNA, tRNA, rRNA, snRNA, etc.) that are necessary for protein synthesis, RNA processing, and gene expression regulation.
3. Translation
Definition
The process of translation involves decoding the genetic information found in a messenger RNA (mRNA) molecule in order to create a particular amino acid sequence in a polypeptide chain.
Similar to transcription, which takes place in the cytoplasm after DNA transcription, it consists of three phases: initiation, elongation, and termination. The elements and processes of DNA translation will be covered in this article.
Components of Translation
Important components include:
• mRNA – template carrying genetic information.
• tRNA – adaptor molecule carrying amino acids.
• Ribosomes – site of protein synthesis.
• Amino acids – building blocks of proteins.
Stages of Translation
Translation occurs in three major steps:
- Initiation
- Elongation
- Termination

Initiation
The small ribosomal subunit binds to mRNA near the start codon (AUG).
Then:
- Initiator tRNA carrying methionine binds to AUG
- Large ribosomal subunit joins to form complete ribosome
The ribosome has three sites:
- A site – aminoacyl site
- P site – peptidyl site
- E site – exit site
Elongation
- The RNA polymerase starts inserting ribonucleotides (A, U, G, C) after initiation to create the RNA strand. The first nucleotide is typically a purine (A or G). Within the transcription bubble, an RNA–DNA hybrid helix (8–9 bases long) develops as the RNA increases.
- The sigma factor is released because it obstructs the RNA exit channel, allowing the core enzyme to keep moving along the DNA template.
- Pyrophosphate is released as each new nucleotide is connected via a phosphodiester linkage.
- Topoisomerases I and II, on the other hand, alleviate the supercoiling that develops in front of the RNA polymerase as it moves.
- The polymerase continues to extend the chain until it encounters a termination signal.
Termination
- Rho-dependent termination: The Rho (ρ) protein, an ATP-dependent RNA helicase, is necessary. Rho binds to a Crich region close to the 3′ end of the RNA transcript and travels along the RNA in the 5′ to 3′ direction. Rho unwinds the RNA-DNA hybrid inside the transcription bubble when it catches up with RNA polymerase at the termination site. The transcription process is terminated when the DNA template, RNA polymerase, and new RNA transcript are released.
- Rho-independent termination: Doesn’t need the Rho protein. An inverted repeat sequence in the DNA generates a GC-rich hairpin loop in the RNA transcript. A series of U residues in RNA (derived from A’s in DNA) follows this loop. The RNA–DNA hybrid is destabilized by the weak A–U base pairings and stable hairpin shape. The outcome is that the DNA template and the RNA transcript are both released as RNA polymerase breaks apart.
Functions of Translation
Translation is essential because it:
- Produces proteins required for cellular functions
- Converts genetic information into functional molecules
- Supports metabolism, structure, and regulation of cells
Proteins produced perform roles such as:
- Enzymatic activity
- Structural support
- Cell signaling Immune defense
Importance of the Central Dogma
The central dogma is fundamental in genetics, biotechnology, and medicine.
Understanding these processes helps in:
- Genetic engineering
- Development of vaccines
- Studying genetic diseases
- Antibiotic development
- Biotechnology research
Modern techniques like PCR, gene cloning, CRISPR, and recombinant DNA technology rely heavily on knowledge of the central dogma.
Conclusion
- The central dogma of molecular biology explains the fundamental flow of genetic information within living cells: DNA → RNA → Protein. Through the processes of replication, transcription, and translation, cells preserve genetic information and convert it into functional molecules necessary for life.
- DNA replication ensures the accurate transmission of genetic information during cell division. Transcription converts DNA sequences into RNA molecules that carry genetic instructions. Translation then interprets these instructions to synthesize proteins that perform vital cellular functions.
- Together, these processes form the foundation of molecular biology and are essential for growth, development, reproduction, and survival of organisms.
- Understanding the central dogma provides insight into how genes control biological traits and how molecular mechanisms regulate life at the cellular level.
Frequently Asked Questions (FAQ)
Q1. What is the central dogma of molecular biology?
The central dogma describes the flow of genetic information in cells from DNA to RNA to protein.
Q2. Who proposed the central dogma?
The concept was proposed by Francis Crick in 1958.
Q3. What are the three steps of the central dogma?
The three main steps are:
- DNA replication
- Transcription
- Translation
Q4. Where does transcription occur?
In eukaryotic cells, transcription occurs in the nucleus, while in prokaryotes, it occurs in the cytoplasm.
Q5. What enzyme is responsible for DNA replication?
The primary enzyme is DNA polymerase, which synthesizes new DNA strands.
Q6. What is a codon?
A codon is a sequence of three nucleotides in mRNA that codes for a specific amino acid.
Q7. What is the role of ribosomes in translation?
Ribosomes serve as the site of protein synthesis, where mRNA is translated into a polypeptide chain.
Reference and Sources
- https://pmc.ncbi.nlm.nih.gov/articles/PMC3685895/
- https://www.genome.gov/genetics-glossary/Central-Dogma
- https://bio.libretexts.org/Bookshelves/Biochemistry/Fundamentals_of_Biochemistry_(Jakubowski_and_Flatt)/03%3A_Unit_III_Information_Pathway/24%3A_DNA_Metabolism/24.01%3A_DNA_Replication
- https://teachmephysiology.com/biochemistry/cell-growth-death/dna-replication/
- https://www.genome.gov/genetics-glossary/Transcription

