SARS: Evolution of a Virus
- In November 2002 a mysterious pneumonia was seen in the Guangdong Province of China, but the first case of this new type of pneumonia was not reported until February 2003.
- Thanks to the ease of global travel, it took only a couple of months for the pneumonia to spread to more than 25 countries in Asia, Europe, and North and South America.
- This newly emergent pneumonia was labeled “severe acute respiratory syndrome” (SARS), and its causative agent was identified as a previously unrecognized coronavirus (CoV), the SARS-CoV.
- Almost 10% of the roughly 8,000 people with SARS died.

- However, once the epidemic was contained, the virus appeared to “die out,” and with the exception of a few mild, sporadic cases in 2004, no additional cases have been identified.
- From where does a newly emergent virus come? And why did this viral disease apparently disappear?
- Coronaviruses are large, enveloped viruses with positive strand RNA genomes. They are known to infect a variety of mammals and birds.
- Researchers suspected that SARS-CoV had “jumped” from its animal host to humans.
- To test this hypothesis, samples of animals at open markets in Guangdong were taken for nucleotide sequencing.
- These studies revealed that cat like animals called masked palm civets (Paguma larvata) harbored variants of the SARS-CoV.
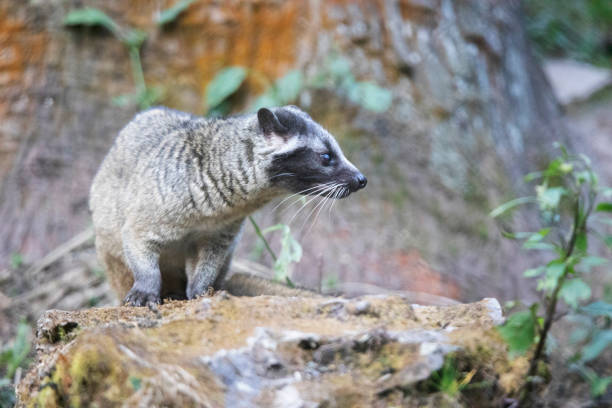

- Although thousands of civets were then slaughtered, further studies failed to find widespread infection of domestic or wild civets.
- In addition, experimental infection of civets with human SARS-CoV strains made these animals ill, making the civet an unlikely candidate for the reservoir species.
- Such a species would be expected to harbor SARS-CoV without symptoms so that it could efficiently spread the virus.
- Bats are reservoir hosts of several zoonotic viruses (viruses spread from animals to people).
- Thus it was perhaps not too surprising when in 2005 two groups of international scientists independently demonstrated that Chinese horseshoe bats (genus Rhinolophus) are the natural reservoir of a SARS like coronavirus.
- When the genomes of the human and bat SARS-CoV are aligned, 92% of the nucleotides are identical.
- More revealing is alignment of the translated amino acid sequences of the proteins encoded by each virus.
- The amino acid sequences are 96 to 100% identical for all proteins except the receptor-binding spike proteins, which are only 64% identical.
- The SARS-CoV spike protein mediates both host cell surface attachment and membrane fusion.
- Thus a mutation of the spike protein allowed the virus to “jump” from bat host cells to those of another species.
- It is not clear if the SARS-CoV was transmitted directly to humans (bats are eaten as a delicacy, and bat feces are a
traditional Asian cure for asthma) or if transmission to humans occurred through infected civets. - The region of the SARS-CoV spike protein that binds to the host receptor, angiotensin-converting enzyme-2 (ACE2), forms a shallow pocket into which ACE2 rests.
- The region of the spike protein that makes this pocket is called the receptor binding domain (RBD).
- Approximately 220 amino acids within the RBD, only four differ between civet and human.
- Two of these amino acids appear to be critical.
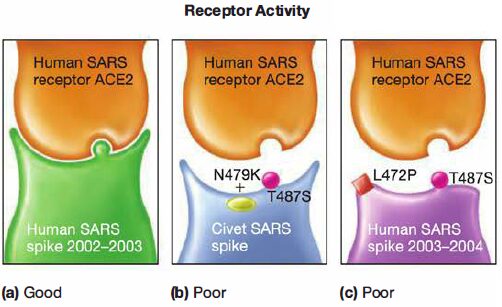
- Compared to the spike RBD in the SARS-CoV that caused the 2002-2003 epidemic, the civet spike has a serine (S) substituted for a threonine (T) at position 487 (T487S) and a lysine (K) at position 479, instead of asparagine (N), N479K.
- This causes a thousand-fold decrease in the capacity of the virus to bind to human ACE2.
- Furthermore, the spike found in SARS-CoV isolated from patients in 2003 and 2004 also has a serine at position 487 as well as a proline (P) for leucine (L) substitution at position 472 (L472P).
- These amino acid substitutions could be responsible for the reduced virulence of the virus found in these more recent infections.
- In other words, these mutations could be the reason the SARS virus appears to have “died out.”
Reference and Sources
- https://www.researchgate.net/publication/7570445_Bats_Are_Natural_Reservoirs_of_SARS-Like_Coronaviruses
- https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5875893/
- https://covid.us.org/2020/06/29/yes-the-coronavirus-really-is-that-bad/
- https://en.wikipedia.org/wiki/History_of_viruses
Also Read:
- Animal and plant viruses, prions, and viroids
- AIDS: Acquired Immune Deficiency Syndrome
- Cider: Production, Extraction, Fermentation and Maturation
- DNA Replication in eukaryotes: Initiation, Elongation and Termination
- Spectroscopy: Introduction, Principles, Types and Applications
- Nosocomial Infection: Introduction, Source, Control and Prevention
- Measurements of microbial growth
- Whole-Genome Shotgun Sequencing: overview, steps and achievements
